EDTA-Resistant, Chemical-Tolerant Ni-Penta™ Agarose-Base Resins for High Selective Purification of His-Tagged Proteins
ProteIndex™ Ni-Penta™ agarose-base resins are advanced, chemical-tolerant immobilized metal ion affinity chromatography (IMAC) media that are precharged with nickel ions. Via a proprietary pentadentate ligand (Figure 1), immobilization of nickel ions on the matrix surface is highly uniform and stable, and offers excellent resistance to a wide range of chemicals, including NaOH, EDTA, reducing agents DTT and β-ME (Refer to Table 1 for the list of tested chemicals).
Owing to this characteristic strong bonding to nickel ions, Ni-Penta™ affinity resins are able to retain high selectivity for polyhistidine in various buffer conditions. They are particularly ideal for purification applications that require the presence of EDTA and reducing agents to maintain integrity and stability of the target proteins. When use Ni-Penta resins to purify the proteins or protein complexes that are difficult to purify with conventional NTA- and IDA- functionalized resins, you now can keep your samples in the most stabilizing conditions during the entire purification procedure. Secondly, Ni-Penta™ resins are well suited for direct capture of polyhistidine-tagged proteins that are secreted into eukaryotic cell culture supernatant (i.e. CHO cells, other mammalian cells, and insect cells). Unlike the conventional IMAC resins, purification with Ni-Penta™ resins generally does not require sample pretreatment to remove EDTA and other agents.
Moreover, the anti-stripping property of Ni-Penta™ contributes to minimal nickel ion leakage from the resin, affording repetitive cycles of stringent clean-in-place (CIP) protocol with 1 M NaOH without the need of regeneration of nickel ions.
Product highlights
 |
|
Nickel ions bind strongly to the functionalized resin surface
The pentadentate ligand on the surface of Ni-Penta™ agarose resin stably immobilizes the nickel ion on the matrix support (Figure 1). The binding between the nickel ion and the agarose support is so strong that it will not be disrupted by high concentration of many commonly used chemicals (see Table 1 for tested chemicals).
 |
Table 1. Chemical compatibility for Chemical-Tolerant Ni-Penta™ Agarose media.
| Chemical Substance | Tested Duration |
| 0.01 M HCl ,0.01 M NaOH | One week |
| 10 mM EDTA, 1 M NaOH, 5 mM DTT, 5 mM TCEP, 20 mM β-mercaptoethanol, 6 M guanidine-HCl | 24 hours |
| 500mM imidazole, 100mM EDTA | 2 hours |
| 30% isopropanol | 20 minutes |
Excellent EDTA-resistance and DTT-tolerance
Chelating agent EDTA, and reducing reagents, such as DTT and β-ME, are able to stabilize proteins from aggregation, and from proteolytic degradation. They are commonly used in cell culture for protein production and sample preparation. When using the traditional NTA- or IDA-based support IMAC resins, however, these reagents will strip off the nickel ions from the functionalized support, making them incompatible with the purification protocol. Removing these reagents during the purification process is necessary, imposing a potential risk on protein integrity and stability or requiring extensive sample handing. The proprietary pentadentate chelating chemistry to immobilize nickel ions on Ni-Penta™ matrix support provides an uncompromised performance in the presence of EDTA and DTT. Shown in Figure 2, the affinity capacity for polyhistidine on Ni-Penta™ Agarose and Ni-Penta™ Agarose 6FF is retained at a comparable level to those with or without 100 mM EDTA and 10 mM DTT.
Therefore, Ni-Penta™ media are well suited for applications of purifying low abundant proteins or challenging proteins from a sample where EDTA, DTT, β-ME, and other stabilizing agents may be necessary for maintaining protein stability and integrity.
Furthermore, Ni-Penta™resins can be used to directly capture polyhistidine-tagged proteins from untreated cell culture supernatant, eliminating the requirement for sample prepretreatment and reducing sample handling (Figure 3).
 |
Figure 2. EDTA- and DTT-compatible affinity of Ni-Penta™ resins to His-tag. Protein loads respectively contain no EDTA/DTT, 100 mM EDTA, and 10 mM DTT. The purified target protein is eluted sequentially with 20 mM and 250 mM imidazole.

Figure 3. Simple purification protocol of of 6xHis-tagged protein secreted into the culture media using Ni-Penta™ resins.
Optimized geometry of nickel ion for better selectivity and higher purity
The surface of Ni-Penta™ resins is modified with a unique, optimized formulation of polydentate-ligand for nickel ion, offering high affinity to polyhistidine and very low background binding with non-specific proteins. In Figure 4, eluates from purification using Ni-Penta™ Agarose resins are analyzed by SDS-PAGE. Shown in the elution profile of a purified His-tagged target protein, the protein eluates from Ni-Penta™ column appeared as a peak with narrower width in the chromatogram, comparable to the eluate profiles when using products from two other suppliers (Figure 5).
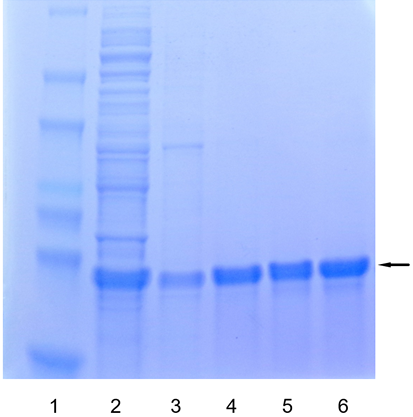 |
Figure 4. Purification of 6xHis-tagged protein on Ni-Penta™ Agarose resins analyzed by SDS-PAGE. Lanes: (1) MW; (2) Raw cell lysate; (3) - (6) Elution fractions.
 |
Figure 5. Purity of polyhistidine-tagged protein on Ni-Penta™ resins. The target protein was loaded on columns packed with Ni-Penta™ Agarose 6FF resin and EDTA-compatible resins from Suppliers G and R, respectively.
Cost-saving by using less imidazole
During purification of His-tagged proteins with conventional Ni-NTA and Ni-IDA based affinity resins, imidazole is generally required to include in the binding and wash buffers to modulate non-specific binding. When using Ni-Penta™ resins, however, most purification protocols require no imidazole in the binding/equilibration buffer, and up to 5 mM in the wash buffer. To elute the target protein from Ni-Penta™ resins, the amount of imidazole may need to be empirically determined. It has been shown that the eluent containing 20 - 40 mM imidazole can be used to elute certain proteins efficiently (Figure 2).
NaOH-tolerant, recharge-free with minimal nickel ion leakage
The highly stable pentadentate-chelating agarose support contributes to minimal nickel ion leakage, and eliminates the need of nickel regeneration even after repetitive Clean-in-Place (CIP) cycles. Shown in Figure 6, dynamic binding capacity for polyhistidine is retained after 100 CIP cycles tested with 0.1 M and 0.5 M NaOH solutions. This ion leakage-proof feature enables a time-efficient, cost-effective, reproducible purification protocol. Furthermore, this property protects your precious protein from toxic nickel as well.
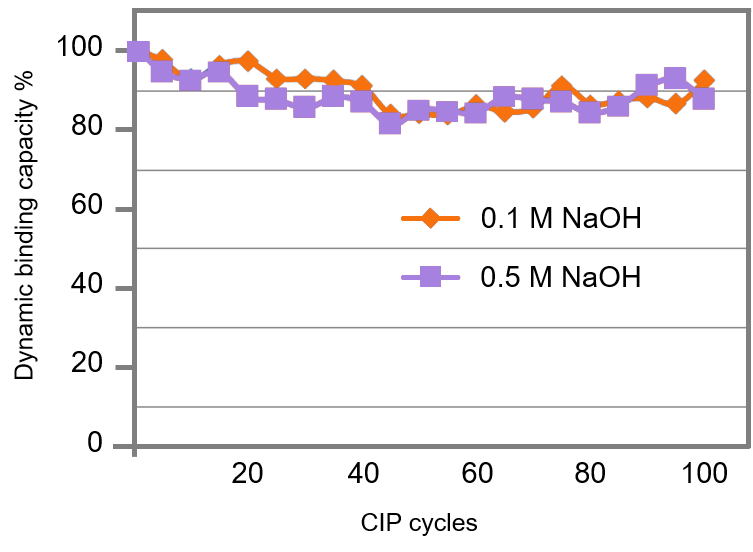 |
Figure 6. Minimal nickel ion leakage from Ni-Penta™ resins after repetitive CIP cycles. Dynamic binding capacity is retained on Ni-Penta™ Agarose 6FF resin after repeated CIP cycles with 0.1 M and 0.5 M NaOH.
Versatile, available in various formats
Marvelgent’s ProteIndex™ Ni-Penta™ resin family includes 4% agarose-based matrix and highly cross-linked 6% agarose-based matrix for various purification needs. They are supplied as settled resins, and are also available as prepacked columns in 1 mL and 5 mL sizes. These products are very flexible and adaptable for screening and preparative purification applications.
To summarize, Ni-Penta resins offer an excellent IMAC alternative to help you obtain your precious proteins in NTA-incompatible conditions. Table 2 summaries some of the specifications of Ni-Penta™ Agarose 6FF, in comparison with similar products that are currently available on the market.

Table 2. Comparison of Ni-Penta™ Agarose 6FF resins with similar products currently available on the market.
Published Mentions
Ni-Penta resins have been referenced in the following publications:
Rewiring photosynthesis: a photosystem I-hydrogenase chimera that makes H2 in vivo. Kanygin A, Milrad Y, Thummala C et al., Energy Environ. Sci., 2020,13, 2903-2914.
The C. difficile toxin B membrane translocation machinery is an evolutionarily conserved protein delivery apparatus. Orrell KE, Mansfield MJ, Doxey AC, Melnyk RA. Nat Commun. 2020 Jan 23;11(1):432. PMID: 31974369
A Unique Protein Self-Assembling Nanoparticle with Significant Advantages in Vaccine Development and Production. DC Carter, B Wright, WG Jerome, JP Rose et al., Journal of Nanomaterials, vol. 2020, Article ID 4297937.
Expression and purification of affinity-tagged variants of the photochemical reaction center from Heliobacterium modesticaldum. Orf GS, Redding KE. Photosynth Res. 2019 Dec;142(3):335-348. PMID: 31542861
Palladin Compensates for the Arp2/3 Complex and Supports Actin Structures during Listeria Infections.Dhanda AS, Vogl AW, Albraiki SE, Otey CA, Beck MR, Guttman JA. MBio. 2018 Apr 10;9(2). PMID: 29636431
Ubiquitinated Proteins Isolated from Tumor Cells Are Efficient Substrates for Antigen. Guangjie Yu, Tarsem Moudgil, Zhihua Cui, Yongbin Mou, Lixin Wang, Bernard A. Fox, and Hong-Ming Hu. J Immunotherapy 2017 Jun;40(5):155-163. PMID: 28368960
Ordering Information
ProteIndex™ Chemical-Tolerant Ni-Penta™ Agarose
| 11-0227-010 | 10 mL settled resins |
| 11-0227-050 | 50 mL settled resins |
| 11-0227-100 | 100 mL settled resins |
ProteIndex™ Chemical-Tolerant Ni-Penta™ Agarose 6 Fast Flow
| 11-0228-010 | 10 mL settled resins |
| 11-0228-050 | 50 mL settled resins |
| 11-0228-100 | 100 mL settled resins |
ProteIndex™ Chemical-Tolerant Ni-Penta™ Agarose 6FF Prepacked Cartridge
| 11-0229-5x1ML | 1 ML Cartridges, Pack of 5 |
| 11-0229-1x5ML | 5 ML Cartridge, Pack of 1 |
| 11-0229-5x5ML | 5 ML Cartridges, Pack of 5 |
Related Products
Tech Note
